alternative splicing
Alternative splicing is a carefully regulated, variable adaptation of the routine RNA modification process of pre-mRNA splicing. Alternative splicing enables a single gene to give rise to multiple versions of a protein. In 1980, a gene called IgM provided the first recognized example of alternative splicing in cells—there were earlier examples in viruses. It has since been demonstrated that cells employ alternative splicing to increase protein diversity toward a variety of biological ends.
A cell typically splices a single transcript in multiple ways to generate an assortment of proteins. Alternatively spliced exons tend to lie between those segments of a gene that encode the functional units, or domains, of a protein (introns). An average mammalian gene possesses eight or nine exons—since most human genes undergoing some form of alternative splicing, virtually all of these exons are candidates for elaborate control.
Alternative splicing depends upon a splice site and nearby enhancer and repressor sequences—short segments of RNA that couple with regulatory proteins. It has been estimated that the splicing of a single exon may be promoted by at least three to seven enhancer sequences. As a result of alternate splicing, mutations that alter a splice site or a nearby regulatory sequence can have subtle effects by shifting the ratio of the resulting proteins without entirely eliminating any form.
Alternative splicing can allow one gene to generate different proteins in different tissues. Many highly specialized brain proteins arise from differential splicing of genes that are also expressed in other tissues. Cells can even modify splicing in response to changing conditions, and not only can alternative splicing tweak the structure of a single protein, but it may also be a means of regulating entire pathways. alternative splicing - click on fig 1 for animation : life cycle of an mRNA ~ click on Quicktime Q :
HHMI Feature Article on Alternative Splicing : Artist's conception of AS
Controlling the Synapse — 49 Proteins at a Time : The Alternative Splicing Website : Alternative Splicing DB (ASDB) : DNA-RNA-ProteinNational Center for Biotechnology Information
A cell typically splices a single transcript in multiple ways to generate an assortment of proteins. Alternatively spliced exons tend to lie between those segments of a gene that encode the functional units, or domains, of a protein (introns). An average mammalian gene possesses eight or nine exons—since most human genes undergoing some form of alternative splicing, virtually all of these exons are candidates for elaborate control.
Alternative splicing depends upon a splice site and nearby enhancer and repressor sequences—short segments of RNA that couple with regulatory proteins. It has been estimated that the splicing of a single exon may be promoted by at least three to seven enhancer sequences. As a result of alternate splicing, mutations that alter a splice site or a nearby regulatory sequence can have subtle effects by shifting the ratio of the resulting proteins without entirely eliminating any form.
Alternative splicing can allow one gene to generate different proteins in different tissues. Many highly specialized brain proteins arise from differential splicing of genes that are also expressed in other tissues. Cells can even modify splicing in response to changing conditions, and not only can alternative splicing tweak the structure of a single protein, but it may also be a means of regulating entire pathways. alternative splicing - click on fig 1 for animation : life cycle of an mRNA ~ click on Quicktime Q :
HHMI Feature Article on Alternative Splicing : Artist's conception of AS
Controlling the Synapse — 49 Proteins at a Time : The Alternative Splicing Website : Alternative Splicing DB (ASDB) : DNA-RNA-ProteinNational Center for Biotechnology Information
base excision repair
Base excision repair describes one form of excision repair in which damaged bases or incorrect bases are excised and replaced by specific enzymes that differ between species. However, the biochemical processes involved in BER are equivalent across species, so bacterial DNA repair functions can operate in eukaryotic cells, and vice versa. Damage is typically the result of deamination, alkylation, hydroxylation, or attack by an oxygen radical, while the incorrect base can be uracil substituted for thymine. Oxidative DNA lesions induced by oxygen free radicals such as superoxide and hydroxyl radicals appear to be repaired predominantly by base excision repair mechanisms. Further, BER is the major DNA repair system involved in removal of various oxidative DNA lesions induced by ionizing radiation - these include abasic sites and modified DNA base and sugar residues.
First, the altered base is excised by a specific DNA glycosylase, which breaks the beta N-glycosidic bond and creates an AP, or abasic site. This site is identical to that generated by spontaneous depyrimidination or depurination. Six DNA glycosylases have been identified in humans – each excises an overlapping subset of either spontaneously formed (such as hypoxanthine), oxidized (such as 8-oxo-7,8-dihydroguanine), alkylated (such as 3-methyladenine), or mismatched (for example, T:G) bases.
Next, the terminal sugar-phosphate is removed by an AP endonuclease (Ape1), leaving a 3’-OH terminal and an abnormal 5'-abasic terminus. The resulting gap is refilled by the 5’-deoxyribose-phosphodiesterase action of a DNA polymerase I (DNA polymerase beta in mammals), then the strands are re-ligated by DNA Ligase I or a complex of XRCC1 and LigIII.
An alternative BER pathway corrects errors involving more than one nucleotide. The Fen1 protein excises the long-patch structure that is produced by DNA polymerase strand displacement. This "long-patch" repair process is divided into two subpathways: a PCNA-stimulated, Pol-beta-directed pathway and a PCNA-dependent, Pol-delta/epsilon -directed pathway.
link to table - human DNA repair genes : diagram>BER
First, the altered base is excised by a specific DNA glycosylase, which breaks the beta N-glycosidic bond and creates an AP, or abasic site. This site is identical to that generated by spontaneous depyrimidination or depurination. Six DNA glycosylases have been identified in humans – each excises an overlapping subset of either spontaneously formed (such as hypoxanthine), oxidized (such as 8-oxo-7,8-dihydroguanine), alkylated (such as 3-methyladenine), or mismatched (for example, T:G) bases.
Next, the terminal sugar-phosphate is removed by an AP endonuclease (Ape1), leaving a 3’-OH terminal and an abnormal 5'-abasic terminus. The resulting gap is refilled by the 5’-deoxyribose-phosphodiesterase action of a DNA polymerase I (DNA polymerase beta in mammals), then the strands are re-ligated by DNA Ligase I or a complex of XRCC1 and LigIII.
An alternative BER pathway corrects errors involving more than one nucleotide. The Fen1 protein excises the long-patch structure that is produced by DNA polymerase strand displacement. This "long-patch" repair process is divided into two subpathways: a PCNA-stimulated, Pol-beta-directed pathway and a PCNA-dependent, Pol-delta/epsilon -directed pathway.
link to table - human DNA repair genes : diagram>BER
cis versus trans-acting factors
Most often, signal elements act only on the intramolecular nucleotide sequence to which they are attached, and they are said to act "in cis". Intron removal in eukaryotes involves cis-splicing. Interaction with signal factors -- usually protein molecules -- turns signal elements on or off.
When protein factors are free to diffuse within the cell they can act on target elements that may not be derived from the same genome segment. Protein factors capable of acting upon other intermolecular genome segments are called "trans-acting factors".
One form of trans-splicing is the 'spliced leader' type, which is primarily found in protozoans (e.g. trypanosomes) and in lower invertebrates such as nematodes. This results in the addition of a capped, noncoding, spliced leader sequence to the 5' end of mRNAs.
Another form of trans-splicing is the 'discontinuous group II intron' type that occurs in plant/algal chloroplasts and plant mitochondria. This results in the joining of two independently transcribed coding sequences. Both spliced-leader and discontinuous group II intron trans-splicing are mechanistically similar to conventional nuclear pre-mRNA cis-splicing. Trans-splicing also occurs in mammalian cells, just as cis-splicing occurs in trypanosomes. It has been suggested that both trans- and cis-splicing are ancient acquisitions of the eukaryotic cell. (Abstract)
When protein factors are free to diffuse within the cell they can act on target elements that may not be derived from the same genome segment. Protein factors capable of acting upon other intermolecular genome segments are called "trans-acting factors".
One form of trans-splicing is the 'spliced leader' type, which is primarily found in protozoans (e.g. trypanosomes) and in lower invertebrates such as nematodes. This results in the addition of a capped, noncoding, spliced leader sequence to the 5' end of mRNAs.
Another form of trans-splicing is the 'discontinuous group II intron' type that occurs in plant/algal chloroplasts and plant mitochondria. This results in the joining of two independently transcribed coding sequences. Both spliced-leader and discontinuous group II intron trans-splicing are mechanistically similar to conventional nuclear pre-mRNA cis-splicing. Trans-splicing also occurs in mammalian cells, just as cis-splicing occurs in trypanosomes. It has been suggested that both trans- and cis-splicing are ancient acquisitions of the eukaryotic cell. (Abstract)
capping
Capping is a form of RNA processing in which the 5’ end of the nascent pre-mRNA is capped with a 7-methyl guanosine nucleotide, 7-methylguanylate. Capping occurs shortly after initiation of transcription.
The 5' cap is retained in mature mRNAs. Capping is required to protect the RNA transcript from degradation. It plays an important role in mRNA transport to the cytoplasm and in the initiation of protein synthesis (translation) . life cycle of an mRNA ~ click on Quicktime Q
NCBI Molecular Cell Biology
Post-transcriptional Processing of RNAs
The 5' cap is retained in mature mRNAs. Capping is required to protect the RNA transcript from degradation. It plays an important role in mRNA transport to the cytoplasm and in the initiation of protein synthesis (translation) . life cycle of an mRNA ~ click on Quicktime Q
NCBI Molecular Cell Biology
Post-transcriptional Processing of RNAs
DNA

DNA is deoxyribonucleic acid, the template for genetic instructions. DNA undergoes replication into complementary strands of DNA for reproduction, and transcription into RNA for the production of proteins. The nucleotide bases, adenine, thymine, cytosine, and guanine are coupled inside the double helix, while the ribose backbone curves around the edge.
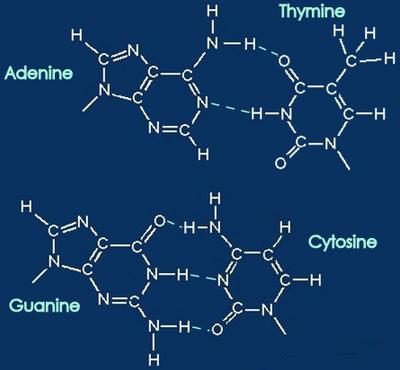 Above - nucleobases form complementary pairs through hydrogen bonding. Adenine and thymine are coupled (broken lines), and cytosine couples with guanine. Below left - stick model with hydrogen bonds as broken lines. (click to enlarge image)
Above - nucleobases form complementary pairs through hydrogen bonding. Adenine and thymine are coupled (broken lines), and cytosine couples with guanine. Below left - stick model with hydrogen bonds as broken lines. (click to enlarge image)
More at:
http://www.dnaftb.org/dnaftb/
http://www.dnalc.org/home.html
http://www.pbs.org/wgbh/aso/tryit/dna/
DNA polymerases
MOLECULAR BIOLOGY: ON DNA POLYMERASES: "Evolution has produced a number of different types of DNA polymerase, but they all have a similar overall three-dimensional shape that has been likened to a right hand, with palm, finger and thumb domains [1]. Polymerases of the A and B families, such as polymerase d, replicate the bulk of genomic DNA during the cell cycle and have been streamlined for processivity and accuracy. These enzymes fit the DNA substrate tightly into their active site, where the replicating base pair is enclosed by the finger domain [2-4]. The mobility of the finger domain underlies a so-called 'induced-fit' mechanism for checking the fidelity of replication: only when an incoming nucleotide forms a perfect Watson-Crick pair with the template base can the fingers close and induce an active conformation of the polymerase. If, nevertheless, an erroneous nucleotide happens to be incorporated, the polymerase responds with conformational distortions of its active center. These induce replication pausing and translocation of the primer terminus towards the intrinsic 'proofreading' exonuclease activity, which removes the mispairing base so that synthesis can resume [5]." O. Fleck and P. Schär (Current Biology 2004 14:R389)
5 Carver, T.E.Jr. , Hochstrasser, R.A., and Millar, D.P. (1994). Proofreading DNA: recognition of aberrant DNA termini by the Klenow fragment of DNA polymerase I. Proc. Natl. Acad. Sci. USA 91, 10670-10674
5 Carver, T.E.Jr. , Hochstrasser, R.A., and Millar, D.P. (1994). Proofreading DNA: recognition of aberrant DNA termini by the Klenow fragment of DNA polymerase I. Proc. Natl. Acad. Sci. USA 91, 10670-10674
DNA damage by ROS
Measurement of Oxidative DNA Damage and Repair in the Aging and Diseased Human Brain Using Liquid and Gas Chromatography/Mass Spectrometry with Isotope Dilution: "There is accumulating evidence that reactive oxygen species (ROS) play an important role in aging and neurodegenerative disease. Genomic DNA appears to be a particularly important target for ROS, since human patients and knockout mice lacking the capacity to repair certain types of [sic] oxidative DNA damage experience neurodegeneration and show evidence of premature aging."
DNA repair
dNTP: "In eukaryotes, DNA damage elicits a multifaceted response that includes cell cycle arrest, transcriptional activation of DNA repair genes, and, in multicellular organisms, apoptosis."
Damage to DNA can be caused by mutations such as replication errors, incorporation of mismatched nucleotides (substitution errors -- transitions and transversions) DNA damage can also result from unintentional and intentional environmental stimuli such as oxygen radicals, hydroxyl radicals, ionizing or ultraviolet radiation, toxins, alkylating agents, and chemotherapy agents, particularly anti-cancer drugs.
Damaged bases can be corrected by simple in situ chemical reversal of the defect, but excision-repair processes predominate. These important DNA repair mechanisms take advantage of the fact that DNA is double-stranded and that complementary sequences should match on both strands. So, if damage is confined to one strand, the damage can be accurately repaired by excision and replacement with DNA synthesized with the undamaged complementary strand acting as template. All organisms, prokaryotic and eukaryotic, utilize at least three enzymatic excision-repair mechanisms: base excision repair, mismatch repair, and nucleotide excision repair.
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMESThe DNA-repair enzymes have the capability of searching through vast tracts of DNA to uncover subtle structural anomalies. The human repair enzyme 8-oxoguanine glycosylase (hOGG1) efficiently removes 8-oxoguanine (oxoG), a damaged guanine (G) base containing an extra oxygen atom, while it ignores undamaged bases. The structure of hOGG1 bound to undamaged DNA, reveals a unique strategy that permits faithful removal of damaged bases which do 'fit' into the oxoG slot at the enzyme's active site, while normal G bases do not.
Anders Jahres medisinske priser: modified: "To date, at least six distinct pathways of DNA repair have been discovered, comprising nucleotide excision repair , base excision repair , mismatch repair , repair by recombination (homologous and nonhomologous end rejoining), damage tolerance pathways (polymerase bypass) and different damage reversal mechanisms, involving close to 200 genes in human cells. "
PLoS Biology: Three New Phases of Repairing DNA Damage in E. coli: "E. coli SOS response has been used to study DNA repair for decades, and a great deal is known about how the more than 30 genes involved in the response function. Two proteins figure prominently in this response. The LexA protein acts as a repressor and inhibits the expression of SOS genes under normal conditions; in the event of DNA damage, the protein RecA inactivates the LexA repressor by enhancing its autocleavage into two fragments, which initiates the SOS response. "
Discovery Of Why Some DNA Repair Fails: Significant For Huntington's Disease And Colon Cancer: "Dr. McMurray's group studied a specific mismatch repair protein Msh2-Msh3 and found a paradox: Instead of helping repair DNA damage, under certain conditions, Msh2-Msh3 was actually harming the cell. Msh2-Msh3 did this when it arrived at the wrong place at the wrong time and bound to a specific portion of DNA (CAG-hairpin). This accident of binding at the CAG-hairpin altered the biochemical activity of Msh2-Msh3. This change in biochemical activity, in turn, promoted DNA expansion -- rather than repair -- and changed the function of Msh2-Msh3 from friend of DNA to foe by allowing damaged DNA to go unrepaired. Without DNA repair, mutations accumulate that lead to disease."
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMES: "hOGG1 makes extensive contacts with the orphaned cytosine base, which ensures that oxoG is removed only when in the appropriate base-pairing context. Although extensive biophysical and structural studies intimate that there are general features of damaged bases that signal their presence to repair enzymes, the steps involved in finding damaged bases in a sea of normal ones are still unclear. Most mechanisms invoke the enzyme sliding or hopping along the DNA duplex until a damaged site is detected. A particularly intriguing question is whether normal bases are also extruded from the helix during the search process."
Damage to DNA can be caused by mutations such as replication errors, incorporation of mismatched nucleotides (substitution errors -- transitions and transversions) DNA damage can also result from unintentional and intentional environmental stimuli such as oxygen radicals, hydroxyl radicals, ionizing or ultraviolet radiation, toxins, alkylating agents, and chemotherapy agents, particularly anti-cancer drugs.
Damaged bases can be corrected by simple in situ chemical reversal of the defect, but excision-repair processes predominate. These important DNA repair mechanisms take advantage of the fact that DNA is double-stranded and that complementary sequences should match on both strands. So, if damage is confined to one strand, the damage can be accurately repaired by excision and replacement with DNA synthesized with the undamaged complementary strand acting as template. All organisms, prokaryotic and eukaryotic, utilize at least three enzymatic excision-repair mechanisms: base excision repair, mismatch repair, and nucleotide excision repair.
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMESThe DNA-repair enzymes have the capability of searching through vast tracts of DNA to uncover subtle structural anomalies. The human repair enzyme 8-oxoguanine glycosylase (hOGG1) efficiently removes 8-oxoguanine (oxoG), a damaged guanine (G) base containing an extra oxygen atom, while it ignores undamaged bases. The structure of hOGG1 bound to undamaged DNA, reveals a unique strategy that permits faithful removal of damaged bases which do 'fit' into the oxoG slot at the enzyme's active site, while normal G bases do not.
Anders Jahres medisinske priser: modified: "To date, at least six distinct pathways of DNA repair have been discovered, comprising nucleotide excision repair , base excision repair , mismatch repair , repair by recombination (homologous and nonhomologous end rejoining), damage tolerance pathways (polymerase bypass) and different damage reversal mechanisms, involving close to 200 genes in human cells. "
PLoS Biology: Three New Phases of Repairing DNA Damage in E. coli: "E. coli SOS response has been used to study DNA repair for decades, and a great deal is known about how the more than 30 genes involved in the response function. Two proteins figure prominently in this response. The LexA protein acts as a repressor and inhibits the expression of SOS genes under normal conditions; in the event of DNA damage, the protein RecA inactivates the LexA repressor by enhancing its autocleavage into two fragments, which initiates the SOS response. "
Discovery Of Why Some DNA Repair Fails: Significant For Huntington's Disease And Colon Cancer: "Dr. McMurray's group studied a specific mismatch repair protein Msh2-Msh3 and found a paradox: Instead of helping repair DNA damage, under certain conditions, Msh2-Msh3 was actually harming the cell. Msh2-Msh3 did this when it arrived at the wrong place at the wrong time and bound to a specific portion of DNA (CAG-hairpin). This accident of binding at the CAG-hairpin altered the biochemical activity of Msh2-Msh3. This change in biochemical activity, in turn, promoted DNA expansion -- rather than repair -- and changed the function of Msh2-Msh3 from friend of DNA to foe by allowing damaged DNA to go unrepaired. Without DNA repair, mutations accumulate that lead to disease."
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMES: "hOGG1 makes extensive contacts with the orphaned cytosine base, which ensures that oxoG is removed only when in the appropriate base-pairing context. Although extensive biophysical and structural studies intimate that there are general features of damaged bases that signal their presence to repair enzymes, the steps involved in finding damaged bases in a sea of normal ones are still unclear. Most mechanisms invoke the enzyme sliding or hopping along the DNA duplex until a damaged site is detected. A particularly intriguing question is whether normal bases are also extruded from the helix during the search process."
targeted genetic repair
Targeted genetic repair: an emerging approach to genetic therapy -- Sullenger 112 (3): 310 -- Journal of Clinical Investigation: "Targeted gene repair is a powerful yet controversial technique developed to direct base changes in chromosomal genes, while RNA repair is an emerging strategy to alter the coding content of messenger RNAs.
Genetic repair strategies may have significant therapeutic and safety advantages over traditional gene therapy approaches for the treatment of many genetic disorders. Firstly, because the mutant genetic instructions are directly repaired, the corrected RNAs and/or DNAs will be maintained in their native sequence context and be regulated by their endogenous regulatory machinery. Secondly, in the instance where the mutant gene encodes a deleterious or dominant-negative mutant protein, repair of the mutant should simultaneously engender the regulated production of the wild-type protein while eliminating or reducing expression of the deleterious gene product. Finally, genetic repair strategies attempt to repair defective instructions in a site-specific manner. Therefore, once adequately developed, these strategies will result in less random mutagenesis of the genome and lead to fewer mutagenic side effects than do methods that randomly insert genes into the genome. "
J. Clin. Invest. 112:310-311 (2003). doi:10.1172/JCI200319419. Perspective : Targeted genetic repair: an emerging approach to genetic therapy
Related Free Full Text articles:
Hacein-Bey-Abina, S. et al.2003. A serious adverse event after successful gene therapy for X-linked severe combined immunodeficiency. N. Engl. J. Med. 348:193-194.[Free Full Text]
Long, M.B., Jones, J.P. III, Sullenger, B.A., and Byun, J. 2003. Ribozyme-mediated revision of RNA and DNA. J. Clin. Invest. 112:312-318. doi:10.1172/JCI200319386.[Free Full Text]
Lan, N. et al.1998. Ribozyme-mediated repair of sickle ß-globin mRNAs in erythrocyte precursors. Science. 280:1593-1596.[Abstract/Free Full Text]
Watanabe, T., and Sullenger, B.A. 2000. Induction of wild-type p53 activity in human cancer cells by ribozymes that repair mutant p53 transcripts. Proc. Natl. Acad. Sci. U. S. A. 97:8490-8494.[Abstract/Free Full Text]
Rogers, C.S., Vanoye, C.G., Sullenger, B.A., and George, A.L. 2002. Functional repair of a mutant chloride channel using a trans-splicing ribozyme. J. Clin. Invest. 110:1783-1789. doi:10.1172/JCI200216481.[Abstract/Free Full Text]
Broitman, S., Amosova, O., Dolinnaya, N.G., and Fresco, J.R. 1999. Repairing the sickle cell mutation. I. Specific covalent binding of a photoreactive third strand to the mutated base pair. J. Biol. Chem. 274:21763-21768.[Abstract/Free Full Text]
Vasquez, K.M., Narayanan, L., and Glazer, P.M. 2000. Specific mutations induced by triplex-forming oligonucleotides in mice. Science. 290:530-533.[Abstract/Free Full Text]
Genetic repair strategies may have significant therapeutic and safety advantages over traditional gene therapy approaches for the treatment of many genetic disorders. Firstly, because the mutant genetic instructions are directly repaired, the corrected RNAs and/or DNAs will be maintained in their native sequence context and be regulated by their endogenous regulatory machinery. Secondly, in the instance where the mutant gene encodes a deleterious or dominant-negative mutant protein, repair of the mutant should simultaneously engender the regulated production of the wild-type protein while eliminating or reducing expression of the deleterious gene product. Finally, genetic repair strategies attempt to repair defective instructions in a site-specific manner. Therefore, once adequately developed, these strategies will result in less random mutagenesis of the genome and lead to fewer mutagenic side effects than do methods that randomly insert genes into the genome. "
J. Clin. Invest. 112:310-311 (2003). doi:10.1172/JCI200319419. Perspective : Targeted genetic repair: an emerging approach to genetic therapy
Related Free Full Text articles:
Hacein-Bey-Abina, S. et al.2003. A serious adverse event after successful gene therapy for X-linked severe combined immunodeficiency. N. Engl. J. Med. 348:193-194.[Free Full Text]
Long, M.B., Jones, J.P. III, Sullenger, B.A., and Byun, J. 2003. Ribozyme-mediated revision of RNA and DNA. J. Clin. Invest. 112:312-318. doi:10.1172/JCI200319386.[Free Full Text]
Lan, N. et al.1998. Ribozyme-mediated repair of sickle ß-globin mRNAs in erythrocyte precursors. Science. 280:1593-1596.[Abstract/Free Full Text]
Watanabe, T., and Sullenger, B.A. 2000. Induction of wild-type p53 activity in human cancer cells by ribozymes that repair mutant p53 transcripts. Proc. Natl. Acad. Sci. U. S. A. 97:8490-8494.[Abstract/Free Full Text]
Rogers, C.S., Vanoye, C.G., Sullenger, B.A., and George, A.L. 2002. Functional repair of a mutant chloride channel using a trans-splicing ribozyme. J. Clin. Invest. 110:1783-1789. doi:10.1172/JCI200216481.[Abstract/Free Full Text]
Broitman, S., Amosova, O., Dolinnaya, N.G., and Fresco, J.R. 1999. Repairing the sickle cell mutation. I. Specific covalent binding of a photoreactive third strand to the mutated base pair. J. Biol. Chem. 274:21763-21768.[Abstract/Free Full Text]
Vasquez, K.M., Narayanan, L., and Glazer, P.M. 2000. Specific mutations induced by triplex-forming oligonucleotides in mice. Science. 290:530-533.[Abstract/Free Full Text]
double strand breaks
MRC Radiation and Genome Stability Unit: "DNA double-strand breaks (DSBs) are potentially lethal, recombinogenic lesions which can result from exposure to DNA damaging agents such as ionising radiation or endogenous events such as collapsed replication forks. Failure to correctly repair a DSB can result in cell death or tumorigenesis. Indeed many cancer genes have been found to function in DSB metabolism. Eukaryotic cells mount a coordinated response to DSBs which includes cell cycle arrest, DNA repair and transcriptional stress responses. These responses are controlled by the DNA integrity checkpoint, DNA repair and stress-activated MAP kinase pathways, respectively. "
enhancer
An enhancer is a short DNA sequence that increases the level of expression of another gene, that is, the enhancer up-regulates transcription of genes within the regulated gene-cluster. Specific trans-acting, transcription factors bind to the enhancer to bring about the increase in transcription rate -- recruiting the initiation complex proteins, or stabilizing the initiation complex.
Because of looping of the DNA strands, there may be a separation of several thousand base pairs between the enhancer and initiator gene (start site). However, the enhancer and its regulated gene are located on the same chromosome.
The enhancer segment may be situated upstream or downstream of the enhanced gene, and its orientation is not fixed – that is, the enhancer’s sequence may be reversed without altering its function. Enhancer segments may be excised and repositioned without interrupting their regulatory function. Enhancers may occur within introns. Enhancers cause the opposite effect to that of silencers which repress transcription.
Because of looping of the DNA strands, there may be a separation of several thousand base pairs between the enhancer and initiator gene (start site). However, the enhancer and its regulated gene are located on the same chromosome.
The enhancer segment may be situated upstream or downstream of the enhanced gene, and its orientation is not fixed – that is, the enhancer’s sequence may be reversed without altering its function. Enhancer segments may be excised and repositioned without interrupting their regulatory function. Enhancers may occur within introns. Enhancers cause the opposite effect to that of silencers which repress transcription.
epigenetics
Defining epigenetic states through chromatin and RNA - Nature Genetics: "The term 'epigenetics' is used to describe heritable changes in genome function that occur without a change in DNA sequence. As such, epigenetics lies at the heart of the cellular memory crucial for development and provides an important avenue for sustained response to environmental stimuli."
exon
Exons are “the sequences of the primary RNA transcript (or the DNA that encodes them) that exit the nucleus as part of a messenger RNA molecule. In the primary transcript neighbouring exons are separated by introns.”
In other words, exons are those sections of DNA within a gene that are not spliced out from the transcribed precursor mRNA and that are retained in the final messenger RNA (mRNA) molecule. For many genes, each exon contains part of the open reading frame (ORF) that codes for a specific portion of the complete protein. However, the term exon is often misused to refer only to coding sequences for the final protein. This is inaccurate since many noncoding exons are known in human genes (Entrez).
The term "exon" was coined by Walter Gilbert in 1978. Gilbert shared the 1980 Nobel Prize in Chemistry with Paul Berg and Frederick Sanger.
The Exon-Intron Database
In other words, exons are those sections of DNA within a gene that are not spliced out from the transcribed precursor mRNA and that are retained in the final messenger RNA (mRNA) molecule. For many genes, each exon contains part of the open reading frame (ORF) that codes for a specific portion of the complete protein. However, the term exon is often misused to refer only to coding sequences for the final protein. This is inaccurate since many noncoding exons are known in human genes (Entrez).
The term "exon" was coined by Walter Gilbert in 1978. Gilbert shared the 1980 Nobel Prize in Chemistry with Paul Berg and Frederick Sanger.
The Exon-Intron Database
gene
The genes are those portions of the genome that code for the production of proteins. Typically protein coding segments are found in the open reading frames of introns, though alternative splicing and other epigenetic modifications account for much of the complexity of the genome.
gene regulation
Gene regulation mechanisms in eukaryotes, which possess a nuclear membrane, differ from those in prokaryotes. Because prokaryotes lack a nuclear membrane, simultaneous translation of a gene may commence before transcription is complete.
In eukaryotes mechanisms for control of gene expression:
- Most commonly affect the rate of transcription.
- Some alter the rate of RNA processing within the nucleus.
- Some affect the stability and degradation of RNA molecules (nonsense-mediated decay, nonstop decay).
- Some control the efficiency of ribosomal translation into ribosomal polypeptides and proteins.
- Some allow for alternative splicing, which generates different proteins from the same archival DNA template.
- Epigenetic mechanisms modify mRNAs.
genome
The term genome refers to the complete hereditary information of an organism (archival DNA or RNA for some viruses). The genome includes both the genes (coding-sequences, domains) and the non-coding sequences – both exons, which include open reading frames, and introns. Similarly, the term proteome refers to an organism’s collection of proteins.
The genome possesses:
- Exonal segments of DNA whose sequence encodes the pre-mRNA, and ultimately polypeptide and protein sequences.
- Intronal segments that are excised by pre-mRNA splicing before transport of mature mRNA through nuclear pores to the cytoplasm where ribosomal translation into ribosomal polypeptides and proteins occurs.
- A start site for transcription, the initiator gene.
- Promoters, both a basal or core promoter located within about 40 bp of the start site, and an upstream promoter, which may extend over as many as 200 bp farther upstream.
- Enhancers.
- Insulators.
- Silencers.
helicases
"Since the discovery of the 'DNA unwinding enzyme,' it has become clear that helicases participate in virtually all cellular processes involving nucleic acids. Helicases are found in all three kingdoms of life and are extremely numerous: 1-2 percent of eukaryotic genes are helicases. Several severe human genetic diseases have been linked to mutations in helicases. . . The most fundamental activity for all helicases is translocation, the ability to move along nucleic acids. Translocation is powered by ATP hydrolysis; hence helicases are motor proteins. Many helicases function as a part of large macromolecular complexes. An example is chromatin remodeling, which regulates gene expression by controlling the DNA access in chromatin. Helicases are the central ATP-powered engines that drive the translocation of the chromatin-remodeling complexes along the DNA. . . Ha's lab measures FRET (fluorescence resonance energy transfer) between various sites on the protein and on the DNA to build dynamic structural models of the protein-DNA complex during translocation. "
HHMI News: DNA Enzyme Shows Unexpected Acrobatic Flair: "Helicase Protein: A helicase protein moving rapidly on a highly flexible single-stranded DNA track. Repetitive movement on the DNA may keep it clear of potentially toxic proteins. Watch Animation at Helicase 8KB Flash Animation(requires Flash Player) "
HHMI News: DNA Enzyme Shows Unexpected Acrobatic Flair: "Helicase Protein: A helicase protein moving rapidly on a highly flexible single-stranded DNA track. Repetitive movement on the DNA may keep it clear of potentially toxic proteins. Watch Animation at Helicase 8KB Flash Animation(requires Flash Player) "
heterochromatin
MOLECULAR BIOLOGY: CHROMATIN DNA PACKAGING AND GENE SILENCING
Cytological studies have demonstrated that much of the repetitious DNA is packaged in a condensed form referred to as heterochromatin. This packaging in the nucleosome limits transcription by rendering DNA segments inaccessible.
The packaged mode of heterochromatin is epigenetically inherited, in that the packaging state is typically maintained after replication and mitosis, independent of the underlying DNA sequence. This property implies a that a biochemical mark exists together with cellular machinery that can recognize and maintain the mark locally.
The heterochromatin of all eukaryotes is characterized by histone hypoacetylation, and by methylation of histone H3 on lysine 9 in higher eukaryotes. Some single-celled eukaryotes such as Saccharomyces lack methylation of H3. Heterochromatin Protein 1 (HP1) binds H3 methylated at lysine 9 (H3-mK9). HP1 is a highly conserved protein that directly associated with pericentric heterochromatin.
A Unified Mode of Epigenetic Gene Silencing RNA Meets Polycomb Group Proteins A Unified Mode of Epigenetic Gene Silencing:
Article Abstract: "Recently, an essential role for RNA in the epigenetic silencing of genes packaged within heterochromatin in animals has been recognized. The RNA appears to be involved in targeting chromatin remodeling activity to specific loci and in later maintaining the repressed state of the gene. Epigenetic silencing of Hox cluster genes by the Polycomb group proteins also involves the formation of a stably inherited repressive chromatin structure. Recent studies of the C. elegans PcG gene sop-2 revealed an evolutionarily conserved property of PcG proteins in the binding of RNA, suggesting an important role for RNA in PcG-mediated Hox gene repression."
Cytological studies have demonstrated that much of the repetitious DNA is packaged in a condensed form referred to as heterochromatin. This packaging in the nucleosome limits transcription by rendering DNA segments inaccessible.
The packaged mode of heterochromatin is epigenetically inherited, in that the packaging state is typically maintained after replication and mitosis, independent of the underlying DNA sequence. This property implies a that a biochemical mark exists together with cellular machinery that can recognize and maintain the mark locally.
The heterochromatin of all eukaryotes is characterized by histone hypoacetylation, and by methylation of histone H3 on lysine 9 in higher eukaryotes. Some single-celled eukaryotes such as Saccharomyces lack methylation of H3. Heterochromatin Protein 1 (HP1) binds H3 methylated at lysine 9 (H3-mK9). HP1 is a highly conserved protein that directly associated with pericentric heterochromatin.
A Unified Mode of Epigenetic Gene Silencing RNA Meets Polycomb Group Proteins A Unified Mode of Epigenetic Gene Silencing:
Article Abstract: "Recently, an essential role for RNA in the epigenetic silencing of genes packaged within heterochromatin in animals has been recognized. The RNA appears to be involved in targeting chromatin remodeling activity to specific loci and in later maintaining the repressed state of the gene. Epigenetic silencing of Hox cluster genes by the Polycomb group proteins also involves the formation of a stably inherited repressive chromatin structure. Recent studies of the C. elegans PcG gene sop-2 revealed an evolutionarily conserved property of PcG proteins in the binding of RNA, suggesting an important role for RNA in PcG-mediated Hox gene repression."
intron
Because the exon segments of archival DNA are transcribed into messenger RNAs and translated into proteins, the earlier view of genes held that introns were solely ‘junk DNA’ or DNA 'deserts' because they do not contain ORFs to code for proteins. However, it has recently been recognized that some exons code for micro RNAs and represent a source of epigenetic coding.
Some segments that were formerly designated introns contain information for protein, and others code for RNA products and are thus not "junk" DNA. Other introns posses translatable nucleotide sequences that, in the absence of splicing, can generate production of novel peptides (maturases) fused to the peptide encoded by the N-terminal exons. In fungi, these peptide maturases, appear to function in intron removal. Their encoding in introns results in homeostatic regulation of their production. Maturase genes are interspersed within other genes.
“Group II introns are a novel class of RNAs (ribozymes) best known for their self-splicing reaction. Under certain in vitro conditions, the introns can excise themselves from precursor mRNAs and ligate together their flanking exons, without the aid of protein. The splicing mechanism is essentially identical to splicing of nuclear pre-mRNA introns (pre-mRNA splicing), and this similarity has led to the widespread belief that group II introns were the ancestors of spliceosomal introns, which make up 25% of the human genome.
Some group II introns have a second remarkable property: they encode reverse transcriptase (RT) ORFs and are active mobile elements. Such mobile group II introns can insert into defined sites at high efficiencies (called retrohoming), or can invade unrelated sites at low frequencies (retrotransposition).”
Ribozyme-mediated revision of RNA and DNA -- Long et al. 112 (3): 312 -- Journal of Clinical Investigation: "Group I introns are ribozymes that carry out two transesterification reactions in order to excise themselves from a precursor transcript. "
Some segments that were formerly designated introns contain information for protein, and others code for RNA products and are thus not "junk" DNA. Other introns posses translatable nucleotide sequences that, in the absence of splicing, can generate production of novel peptides (maturases) fused to the peptide encoded by the N-terminal exons. In fungi, these peptide maturases, appear to function in intron removal. Their encoding in introns results in homeostatic regulation of their production. Maturase genes are interspersed within other genes.
“Group II introns are a novel class of RNAs (ribozymes) best known for their self-splicing reaction. Under certain in vitro conditions, the introns can excise themselves from precursor mRNAs and ligate together their flanking exons, without the aid of protein. The splicing mechanism is essentially identical to splicing of nuclear pre-mRNA introns (pre-mRNA splicing), and this similarity has led to the widespread belief that group II introns were the ancestors of spliceosomal introns, which make up 25% of the human genome.
Some group II introns have a second remarkable property: they encode reverse transcriptase (RT) ORFs and are active mobile elements. Such mobile group II introns can insert into defined sites at high efficiencies (called retrohoming), or can invade unrelated sites at low frequencies (retrotransposition).”
Ribozyme-mediated revision of RNA and DNA -- Long et al. 112 (3): 312 -- Journal of Clinical Investigation: "Group I introns are ribozymes that carry out two transesterification reactions in order to excise themselves from a precursor transcript. "
mismatch repair
Most mismatches are caused by replication errors. However, mismatches can also be produced by other mechanisms, such as deamination of 5-methyl cytosine to generate T improperly paired to G. Where the appropriate DNA-N-glycosylase is available, mismatches can also be repaired by base excision repair.
Mismatch repair has been studied most extensively in E. coli, where the proteins MutS, MutL, and MutH initiate the repair process. Newly synthesized strands are not immediately methylated in E.coli. First, MutS recognizes and binds to true mismatches and insertions/deletions of up to 4 nucleotides. Next, MutL binds to and stabilizes this complex of MutS/mismatched strand. The MutS-MutL complex then activates MutH, which locates a nearby methyl group and creates a nick in the newly synthesized strand opposite the methyl group. Excision is accomplished in E.coli by cooperation between the UvrD (Helicase II) protein, which unwinds from the nick toward the mismatch, and a single-strand specific exonuclease of appropriate polarity. Finally, resynthesis by Polymerase III and ligation by a DNA ligase repair the sequence and re-ligate the strands.
Unlike the un-methylated new strands in E.coli, strand-specificity in eukaryotes may be signalled by single-strand nicks. In nascent eukaryotic DNA strands, single-strand breaks occur between Okazaki fragments in the lagging strand and at the 3' end of the leading strand.
Eukaryotes lack homologues of MutH and uvrD, but do possess numerous homologues of MutS and MutL (MSHs 1-6, MLHs 1-3 and PMS 1 or 2). In E.coli, MutS and MutL function as monomers. The homologous eukaryotic proteins function as heterodimers. Human cells also possess two heterodimers of MutS homologues – MutSalpha (MSH2/MSH6), which recognizes single base mismatches and small loops, and MutSbeta (MSH2/MSH3), which recognizes small loops.
In eukaryotic cells, several standard replication proteins are needed for mismatch repair. The "clamp" protein, PCNA is a cofactor for most DNA polymerases and stabilizes the MutS and MutL heterodimers at mismatch sites on DNA. Three MutL homolog dimers are also known – in humans, MLH1/PMS2, MLH1/PMS1, and MLH1/MLH3. Further, just as exonucleases are thought to be important for mismatch repair in prokaryotes, at least two nucleases appear to contribute to mismatch repair in eukaryotic cells – exonuclease 1 and Flap Endonuclease (FEN-1 or DNase IV; Rad27 in S. cerevisiae). PCNA is also required during the later DNA synthesis step of mismatch repair. The DNA synthesis step also requires RPA (the eukaryotic single-stranded DNA-binding protein), Replication factor C (which loads PCNA onto DNA molecules at primer termini) and DNA polymerase delta.
Hereditary non-polyposis colon cancer (HNPCC) is a form of colon cancer frequently associated with defects in the genes encoding MSH2 (about 35% of identified gene-defect cases) and MLH1 (about 60% of identified gene-defect cases). HNPCC is characterized by early age of onset and autosomal dominant inheritance with high penetrance.
Mismatch repair has been studied most extensively in E. coli, where the proteins MutS, MutL, and MutH initiate the repair process. Newly synthesized strands are not immediately methylated in E.coli. First, MutS recognizes and binds to true mismatches and insertions/deletions of up to 4 nucleotides. Next, MutL binds to and stabilizes this complex of MutS/mismatched strand. The MutS-MutL complex then activates MutH, which locates a nearby methyl group and creates a nick in the newly synthesized strand opposite the methyl group. Excision is accomplished in E.coli by cooperation between the UvrD (Helicase II) protein, which unwinds from the nick toward the mismatch, and a single-strand specific exonuclease of appropriate polarity. Finally, resynthesis by Polymerase III and ligation by a DNA ligase repair the sequence and re-ligate the strands.
Unlike the un-methylated new strands in E.coli, strand-specificity in eukaryotes may be signalled by single-strand nicks. In nascent eukaryotic DNA strands, single-strand breaks occur between Okazaki fragments in the lagging strand and at the 3' end of the leading strand.
Eukaryotes lack homologues of MutH and uvrD, but do possess numerous homologues of MutS and MutL (MSHs 1-6, MLHs 1-3 and PMS 1 or 2). In E.coli, MutS and MutL function as monomers. The homologous eukaryotic proteins function as heterodimers. Human cells also possess two heterodimers of MutS homologues – MutSalpha (MSH2/MSH6), which recognizes single base mismatches and small loops, and MutSbeta (MSH2/MSH3), which recognizes small loops.
In eukaryotic cells, several standard replication proteins are needed for mismatch repair. The "clamp" protein, PCNA is a cofactor for most DNA polymerases and stabilizes the MutS and MutL heterodimers at mismatch sites on DNA. Three MutL homolog dimers are also known – in humans, MLH1/PMS2, MLH1/PMS1, and MLH1/MLH3. Further, just as exonucleases are thought to be important for mismatch repair in prokaryotes, at least two nucleases appear to contribute to mismatch repair in eukaryotic cells – exonuclease 1 and Flap Endonuclease (FEN-1 or DNase IV; Rad27 in S. cerevisiae). PCNA is also required during the later DNA synthesis step of mismatch repair. The DNA synthesis step also requires RPA (the eukaryotic single-stranded DNA-binding protein), Replication factor C (which loads PCNA onto DNA molecules at primer termini) and DNA polymerase delta.
Hereditary non-polyposis colon cancer (HNPCC) is a form of colon cancer frequently associated with defects in the genes encoding MSH2 (about 35% of identified gene-defect cases) and MLH1 (about 60% of identified gene-defect cases). HNPCC is characterized by early age of onset and autosomal dominant inheritance with high penetrance.
nonsense-mediated decay
Defusing Dangerous Mutations: Scientists Discover A New Way By Which Cells Control Genetic Errors: Adapted: "Nonsense-Mediated Decay (NMD), is a process by which cells destroy potentially harmful molecules. Both healthy and damaged proteins begin as instructions in genes. Cells transcribe this information and create an mRNA molecule, a template that will be used to create proteins. RNAs usually contain extra bits of code that have to be cut out before they can be used. During this cut-and-paste operation, cells attach a group of molecules called the exon junction complex (EJC) to the RNA. An RNA made from a mutant gene usually has an EJC in the wrong position, which activates NMD and destroys the RNA before it can be used to make flawed proteins. There are at least two kinds of NMD: one requires UPF2 and the other does not.
The presence or absence of UPF2 changes the composition of the EJC, giving it different surfaces to which other molecules attach. This affects the way that another component, called UPF1, fits onto the machine. UPF1 is directly responsible for calling up the NMD machinery. The study shows that UPF1 can be mounted on both EJC types; the final effect is the same – to efficiently destroy faulty RNAs. "
The presence or absence of UPF2 changes the composition of the EJC, giving it different surfaces to which other molecules attach. This affects the way that another component, called UPF1, fits onto the machine. UPF1 is directly responsible for calling up the NMD machinery. The study shows that UPF1 can be mounted on both EJC types; the final effect is the same – to efficiently destroy faulty RNAs. "
nonstop decay
Researchers Discover New Mechanism That Targets And Destroys Abnormal RNA:
Adapted: "Messenger RNA molecules are the genetic templates for proteins. In constructing proteins, the mRNA template is transcribed from DNA genes and transported to the ribosomes -- the cell's protein "factories" that are large complexes of protein and RNA. Given the importance of mRNA as an information-carrying molecule, the machinery that regulates mRNA levels and destroys faulty mRNA is critical in ensuring that errors in the genetic code are not passed on to proteins.
Nonstop decay is mRNA turnover mechanism that has none of the properties of nonsense-mediated decay (NMD), or of normal mRNA turnover in the cell. Nonstop decay shares none of the enzymes required for nonsense-mediated decay. A multi-enzyme complex called the exosome is important for nonstop decay, site of binding of a specific adapter protein called Ski7p occurs."
One percent of genes in both humans and yeast produce mRNAs containing specific sequences that would trigger degradation of the RNA by nonstop decay. Nonstop mRNA transcripts might be important in enabling production of shortened proteins that are needed at specific stages of development. At later stages of development, when these truncated proteins are no longer needed, their mRNA could easily be destroyed by nonstop decay."
The original news release can be found here.
Adapted: "Messenger RNA molecules are the genetic templates for proteins. In constructing proteins, the mRNA template is transcribed from DNA genes and transported to the ribosomes -- the cell's protein "factories" that are large complexes of protein and RNA. Given the importance of mRNA as an information-carrying molecule, the machinery that regulates mRNA levels and destroys faulty mRNA is critical in ensuring that errors in the genetic code are not passed on to proteins.
Nonstop decay is mRNA turnover mechanism that has none of the properties of nonsense-mediated decay (NMD), or of normal mRNA turnover in the cell. Nonstop decay shares none of the enzymes required for nonsense-mediated decay. A multi-enzyme complex called the exosome is important for nonstop decay, site of binding of a specific adapter protein called Ski7p occurs."
One percent of genes in both humans and yeast produce mRNAs containing specific sequences that would trigger degradation of the RNA by nonstop decay. Nonstop mRNA transcripts might be important in enabling production of shortened proteins that are needed at specific stages of development. At later stages of development, when these truncated proteins are no longer needed, their mRNA could easily be destroyed by nonstop decay."
The original news release can be found here.
nucleic acids
A nucleic acid is a large biochemical macromolecule composed of chains of nucleotides coding genetic information. The most common nucleic acids are deoxyribonucleic acid ( DNA ) and ribonucleic acid ( RNA ). Nucleic acids are found in all living cells – karyotes and eukaryotes – and in viruses. Monomeric nucleic acids are called nucleotides – each consists of a nitrogenous heterocyclic base (either a purine or a pyrimidine), a five-ring pentose sugar, and a phosphate group.
nucleosome
MOLECULAR BIOLOGY: CHROMATIN DNA PACKAGING AND GENE SILENCING
The nucleosome is the basic repeat element of chromatin, and consists of 147 base pairs (bp) of DNA wrapped 1.7 times around an octamer of histone proteins (two copies each of the core histones H2A, H2B, H3, and H4).
Nucleosomes are connected by about 20 to 60 bp of linker DNA to form the 10-nm "beads-on-a-string" array. This can be further compacted into a "30-nm" chromatin fiber.
Two classes of model for chromatin have been proposed: (a) the "one-start helix" in which nucleosomes, connected by bent linker DNA, are arranged linearly in a higher order helix; or (b) the "two-start helix" in which nucleosomes, connected by straight linker DNA, zigzag back and forth between two adjacent helical stacks.
To distinguish between these two competing models of higher order chromatin folding, Dorigo and co-workers employed a fully defined in vitro system to generate regular nucleosomal arrays. Analysis of the length of the nucleosome stacks, now connected only by internucleosomal cross-links, revealed a two-start rather than a one-start organization. This interpretation was corroborated by electron microscopy. Thus, local interactions between nucleosomes can drive self-organization into a higher order chromatin fiber. Adapted from: Adone Mohd-Sarip and C. Peter Verrijzer (Science 2004 306:1484)
1. B. Dorigo et al., Science 306, 1571 (2004)
The nucleosome is the basic repeat element of chromatin, and consists of 147 base pairs (bp) of DNA wrapped 1.7 times around an octamer of histone proteins (two copies each of the core histones H2A, H2B, H3, and H4).
Nucleosomes are connected by about 20 to 60 bp of linker DNA to form the 10-nm "beads-on-a-string" array. This can be further compacted into a "30-nm" chromatin fiber.
Two classes of model for chromatin have been proposed: (a) the "one-start helix" in which nucleosomes, connected by bent linker DNA, are arranged linearly in a higher order helix; or (b) the "two-start helix" in which nucleosomes, connected by straight linker DNA, zigzag back and forth between two adjacent helical stacks.
To distinguish between these two competing models of higher order chromatin folding, Dorigo and co-workers employed a fully defined in vitro system to generate regular nucleosomal arrays. Analysis of the length of the nucleosome stacks, now connected only by internucleosomal cross-links, revealed a two-start rather than a one-start organization. This interpretation was corroborated by electron microscopy. Thus, local interactions between nucleosomes can drive self-organization into a higher order chromatin fiber. Adapted from: Adone Mohd-Sarip and C. Peter Verrijzer (Science 2004 306:1484)
1. B. Dorigo et al., Science 306, 1571 (2004)
nucleotide
Single chain (monomeric) nucleic acids are called nucleotides -- each consists of a nitrogenous heterocyclic base (either a purine or a pyrimidine), a pentose sugar, and a phosphate group.
nucleotide excision repair
A nucleotide with abnormal alterations in its chemical properties is termed an elementary damage site (EDS). UV light is a frequent cause of damage to segments of DNA. Nucleotide excision repair is a multi-enzyme excision repair pathway targetted at damaged segments of DNA. Similar mechanisms exist in both prokaryotes and eukaryotes, and the mechanism is highly conserved in eukaryotes. The recognition of DNA damage is less specific in nucleotide excision repair (NER) than in base excision repair (BER), which targets damaged bases. Thus NER repairs a wider range of damage types than BER, irrespective of chromatin structure or the gene expression profile of a particular cell.
NER operates by: (a) recognition of the damaged DNA; (b) excision of an oligonucleotide of 24–32 residues by dual incision of the damaged strand on each side of the lesion; and, as for BER (c) filling in of the resulting gap by DNA polymerase; and (d) ligation of the nick. There is evidence that at least some steps of NER require ATP-dependent chromatin remodeling activities.
NER can operate by two pathways. The first pathway, global genome repair (GGR), acts on DNA lesions across the genome and is transcription-independent. The second NER pathway, transcription coupled repair, is coupled to active transcription and is directed to the transcribed strand of active genes.
clickable diagram - nucleotide excision repair : diagram>nucleotide excision repair : diagram>transition coupled repair : more detail NER defective medical disorders : free full text review article :
NER operates by: (a) recognition of the damaged DNA; (b) excision of an oligonucleotide of 24–32 residues by dual incision of the damaged strand on each side of the lesion; and, as for BER (c) filling in of the resulting gap by DNA polymerase; and (d) ligation of the nick. There is evidence that at least some steps of NER require ATP-dependent chromatin remodeling activities.
NER can operate by two pathways. The first pathway, global genome repair (GGR), acts on DNA lesions across the genome and is transcription-independent. The second NER pathway, transcription coupled repair, is coupled to active transcription and is directed to the transcribed strand of active genes.
clickable diagram - nucleotide excision repair : diagram>nucleotide excision repair : diagram>transition coupled repair : more detail NER defective medical disorders : free full text review article :
nuclear speckles
New Gene Regulation Mechanism Discovered: "Dr. David Spector noticed that under standard growth conditions, a particular population of messenger RNA molecules lingered in the nucleus indefinitely--in structures they call 'nuclear speckles'--and never reached the cytoplasm.
One of Spector's graduate students developed a method for purifying speckles. That allowed the researchers to identify not only the many different protein components of speckles, but also the messenger RNAs that are the basis of the new study, published in the October 21 issue of the journal Cell. The study--spearheaded by Cold Spring Harbor Laboratory postdoctoral fellow Dr. Kannanganattu Prasanth--identified the first such messenger RNA: one transcribed from a mouse gene called mCAT2 that encodes a cell surface receptor.
The scientists learned from the work of others that the mCAT2 receptor is involved in the production of nitric oxide, and that nitric oxide production is stimulated by various stress conditions including wound healing and viral infection. " That told us that when cells are stressed, maybe the atypical messenger RNA is released from the nucleus, exported to the cytoplasm, and translated into protein, thus circumventing the time-consuming process of producing new messenger RNA and providing a rapid response to viral infection or other stresses," says Spector. To test this idea, the researchers mimicked the effect of viral infection by treating cells with interferon.
Sure enough, they discovered that the atypical mCAT2 messenger RNA in the nucleus was rapidly cleaved in response to interferon treatment, and that the protein coding portion of the molecule was then quickly exported to the cytoplasm and translated into protein. diagram.
" This 'cut and run' mechanism is a completely new paradigm of gene regulation, so studying it will keep us busy for a while. But we already suspect that there is going to be a large family of genes regulated in this way," says Spector.""
One of Spector's graduate students developed a method for purifying speckles. That allowed the researchers to identify not only the many different protein components of speckles, but also the messenger RNAs that are the basis of the new study, published in the October 21 issue of the journal Cell. The study--spearheaded by Cold Spring Harbor Laboratory postdoctoral fellow Dr. Kannanganattu Prasanth--identified the first such messenger RNA: one transcribed from a mouse gene called mCAT2 that encodes a cell surface receptor.
The scientists learned from the work of others that the mCAT2 receptor is involved in the production of nitric oxide, and that nitric oxide production is stimulated by various stress conditions including wound healing and viral infection. " That told us that when cells are stressed, maybe the atypical messenger RNA is released from the nucleus, exported to the cytoplasm, and translated into protein, thus circumventing the time-consuming process of producing new messenger RNA and providing a rapid response to viral infection or other stresses," says Spector. To test this idea, the researchers mimicked the effect of viral infection by treating cells with interferon.
Sure enough, they discovered that the atypical mCAT2 messenger RNA in the nucleus was rapidly cleaved in response to interferon treatment, and that the protein coding portion of the molecule was then quickly exported to the cytoplasm and translated into protein. diagram.
" This 'cut and run' mechanism is a completely new paradigm of gene regulation, so studying it will keep us busy for a while. But we already suspect that there is going to be a large family of genes regulated in this way," says Spector.""
open reading frame
Because codons occur in nucleotide triplets, from any point within the genome there are six possible reading frames (three in each direction). Open reading frames (ORFs) are those sequences of DNA or RNA that code for translation at ribosomes into polypeptides or proteins. Within the genome, (ORFs) lie between codons for initiation (start-code sequence) and termination (stop-code sequence).
A DNA open reading frame starts with ATG—coding for Met—in most species, and ends with a stop codon (TAA, TAG, or TGA). In prokaryotes, ORFs are usually the longest sequence without a stop codon. In eukaryotes, long ORFs can continue over non-translatable intron gaps, so spliced mRNA must be employed to determine ORFs. Short ORFs can occur outside genes – within the intron, segments of DNA outside genes that were formerly considered ‘junk’ DNA.
A DNA open reading frame starts with ATG—coding for Met—in most species, and ends with a stop codon (TAA, TAG, or TGA). In prokaryotes, ORFs are usually the longest sequence without a stop codon. In eukaryotes, long ORFs can continue over non-translatable intron gaps, so spliced mRNA must be employed to determine ORFs. Short ORFs can occur outside genes – within the intron, segments of DNA outside genes that were formerly considered ‘junk’ DNA.
oxoG repair
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMES
The agents that cause oxidative damage to DNA include oxygen radicals and ionizing radiation. Oxidation of a guanosine base to form oxoG produces a subtle structural transformation that results in deleterious mutations because DNA polymerases misread oxoG as a thymine (T) during genome replication prior to cell division. The human oxoG repair enzyme (hOGG1) catalyses the excision of oxoG in the first step of base excision repair.
Structural studies of the glycosylase enzymes involved in the repair process reveal common features of damaged-base recognition. These include enzyme-initiated DNA distortion that flips the damaged base out from the DNA double helix for recognition within a base-specific cavity of the enzyme. (OGG1)
The agents that cause oxidative damage to DNA include oxygen radicals and ionizing radiation. Oxidation of a guanosine base to form oxoG produces a subtle structural transformation that results in deleterious mutations because DNA polymerases misread oxoG as a thymine (T) during genome replication prior to cell division. The human oxoG repair enzyme (hOGG1) catalyses the excision of oxoG in the first step of base excision repair.
Structural studies of the glycosylase enzymes involved in the repair process reveal common features of damaged-base recognition. These include enzyme-initiated DNA distortion that flips the damaged base out from the DNA double helix for recognition within a base-specific cavity of the enzyme. (OGG1)
oxidative stress and DNA damage
Oxidative Stress/DNA Damage and DNA Repair: "Oxidative stress is produced in cells by oxygen-derived species resulting from cellular metabolism and from interaction with cells of exogenous sources such as carcinogenic compounds, redox-cycling drugs and ionizing radiations. DNA damage caused by oxygen-derived species including free radicals is the most frequent type encountered by aerobic cells. DNA damage caused by oxygen-derived species including free radicals is the most frequent type encountered by aerobic cells. When this type of damage occurs to DNA, it is called oxidative DNA damage and it can produce a multiplicity of modifications in DNA including base and sugar lesions, strand breaks, DNA-protein cross-links and base-free sites. "
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMESThe agents that cause oxidative damage to DNA include oxygen radicals and ionizing radiation. Oxidation of a guanosine base to form oxoG produces a subtle structural transformation that results in deleterious mutations because DNA polymerases misread oxoG as a thymine (T) during genome replication prior to cell division. The human oxoG repair enzyme (hOGG1) catalyses the excision of oxoG in the first step of base excision repair. Structural studies of the glycosylase enzymes involved in the repair process reveal common features of damaged-base recognition. These include enzyme-initiated DNA distortion that flips the damaged base out from the DNA double helix for recognition within a base-specific cavity of the enzyme. (OGG1)
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMES: "hOGG1 makes extensive contacts with the orphaned cytosine base, which ensures that oxoG is removed only when in the appropriate base-pairing context. Although extensive biophysical and structural studies intimate that there are general features of damaged bases that signal their presence to repair enzymes, the steps involved in finding damaged bases in a sea of normal ones are still unclear. Most mechanisms invoke the enzyme sliding or hopping along the DNA duplex until a damaged site is detected. A particularly intriguing question is whether normal bases are also extruded from the helix during the search process."
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMESThe agents that cause oxidative damage to DNA include oxygen radicals and ionizing radiation. Oxidation of a guanosine base to form oxoG produces a subtle structural transformation that results in deleterious mutations because DNA polymerases misread oxoG as a thymine (T) during genome replication prior to cell division. The human oxoG repair enzyme (hOGG1) catalyses the excision of oxoG in the first step of base excision repair. Structural studies of the glycosylase enzymes involved in the repair process reveal common features of damaged-base recognition. These include enzyme-initiated DNA distortion that flips the damaged base out from the DNA double helix for recognition within a base-specific cavity of the enzyme. (OGG1)
MOLECULAR BIOLOGY: ON DNA-REPAIR ENZYMES: "hOGG1 makes extensive contacts with the orphaned cytosine base, which ensures that oxoG is removed only when in the appropriate base-pairing context. Although extensive biophysical and structural studies intimate that there are general features of damaged bases that signal their presence to repair enzymes, the steps involved in finding damaged bases in a sea of normal ones are still unclear. Most mechanisms invoke the enzyme sliding or hopping along the DNA duplex until a damaged site is detected. A particularly intriguing question is whether normal bases are also extruded from the helix during the search process."
polyadenylation
Polyadenylation is a form of RNA processing in which the 3’ end of the pre-mRNA is cleaved before a stretch of adenosines are added to the end of the molecule. Employment of alternative polyadenylation sites can result in the insertion or deletion of sequences that control the stability of the mRNA, and thus the level of protein expression. The polyA tail is also involved in initiation of translation.
MOLECULAR BIOLOGY: ON TRANSCRIPTION TERMINATION: "A defining feature of mRNAs is a tail composed of a long string of adenosine nucleotides -- the poly(A) tail. This is not encoded by the gene but is added following cleavage of the nascent RNA transcript. Molecular factors that recognize the cleavage site, and cut the RNA, bind to a regulatory region of the transcribing RNA polymerase II called the carboxy-terminal domain (CTD). This interaction is important for recruiting the factors to the nascent transcript. In turn, these factors must be off-loaded from the polymerase onto the RNA at their site of action for transcription to be terminated(4,5). In fact, recognition of the cleavage site has been believed to be the key step in termination -- but the new data(1-3) show that this isn't the whole story."
MOLECULAR BIOLOGY: ON TRANSCRIPTION TERMINATION: "A defining feature of mRNAs is a tail composed of a long string of adenosine nucleotides -- the poly(A) tail. This is not encoded by the gene but is added following cleavage of the nascent RNA transcript. Molecular factors that recognize the cleavage site, and cut the RNA, bind to a regulatory region of the transcribing RNA polymerase II called the carboxy-terminal domain (CTD). This interaction is important for recruiting the factors to the nascent transcript. In turn, these factors must be off-loaded from the polymerase onto the RNA at their site of action for transcription to be terminated(4,5). In fact, recognition of the cleavage site has been believed to be the key step in termination -- but the new data(1-3) show that this isn't the whole story."
RNA processing
Posttranscriptional RNA processing is an important feature of eukaryotic cells.
Pre-mRNA processing takes place in the nucleus. RNA processing events include capping of the 5’ end on the pre-mRNA, pre-mRNA splicing to remove intronic sequences, and polyadenylation of the 3’ end of the pre-mRNA.
Capping of the 5’ end of the nascent pre-mRNA is performed soon after initiation of transcription. Cleavage, pre-mRNA splicing, and polyadenylation usually follow termination of transcription of short primary transcripts with few introns. However, introns often are spliced out of the nascent RNA before transcription of the gene is complete for large genes with multiple introns. diagram - pre-mRNA processing : animation of RNA splicing : animation - life cycle of an mRNA : diagram - spliceosome assembly :
NCBI Molecular Cell Biology
Post-transcriptional Processing of RNAs
Pre-mRNA processing takes place in the nucleus. RNA processing events include capping of the 5’ end on the pre-mRNA, pre-mRNA splicing to remove intronic sequences, and polyadenylation of the 3’ end of the pre-mRNA.
Capping of the 5’ end of the nascent pre-mRNA is performed soon after initiation of transcription. Cleavage, pre-mRNA splicing, and polyadenylation usually follow termination of transcription of short primary transcripts with few introns. However, introns often are spliced out of the nascent RNA before transcription of the gene is complete for large genes with multiple introns. diagram - pre-mRNA processing : animation of RNA splicing : animation - life cycle of an mRNA : diagram - spliceosome assembly :
NCBI Molecular Cell Biology
Post-transcriptional Processing of RNAs
pre-mRNA
Precursor messenger RNAs, or heteronuclear RNAs (hnRNAs) are products of gene transcription found in the cell's nucleus, which contain bases complementary to those of the template DNA strand, including both exons and introns.
Following pre-mRNA processing, which includes capping, pre-mRNA splicing, alternative splicing, and polyadenylation, the mature mRNA is transported through nuclear pores into the cytoplasm.
Following pre-mRNA processing, which includes capping, pre-mRNA splicing, alternative splicing, and polyadenylation, the mature mRNA is transported through nuclear pores into the cytoplasm.
pre-mRNA splicing
Because eukaryotic RNAs are transcribed from intron containing genes, the sequences encoded by the intronic DNA must be removed from the primary transcript prior to the RNAs becoming biologically active. The process of intron removal is called RNA splicing, or pre-mRNA splicing. The intron-exon junctions (splice sites) in the precursor mRNA (pre-mRNA) are recognized by trans-acting factors (prokaryote RNAs are mostly polycistronic). In pre-mRNA splicing the intronic sequences are excised and the exons are ligated to generate the spliced mRNA.
Group I introns occur in nuclear, mitochondrial and chloroplast rRNA genes, group II in mitochondrial and chloroplast mRNA genes. Many of the group I and group II introns are self-splicing in that no additional protein factors are necessary for the intron to be efficiently and accurately excised and the strands reattached.
Group I introns require an external guanosine nucleotide as a cofactor. The 3'-OH of the guanosine nucleotide acts as a nucleophile to attack the 5'-phosphate of the intron's 5' nucleotide. The 3' end of the 5' exon is termed the splice donor site. The 3'-OH at the 3' splice donor end of the 5' exon next attacks the splice acceptor site at the 5' nucleotide of the 3' exon, releasing the intron and covalently attaching the two exons together.
Pre-mRNA processing takes place in the nucleus of eukaryotes, whereas lack of a nuclear membrane in prokaryotes permits initiation of translation while transcription is not yet complete.
Pre-mRNA processing events include capping of the 5’ end on the pre-mRNA, pre-mRNA splicing to remove intronic sequences, and polyadenylation of the 3’ end of the pre-mRNA. animation of RNA splicing requires Flash Player plugin - Download plugin: clickable slide show - spliceosome intron removal :
More in NCBI Molecular Cell Biology on-line text.
Group I introns occur in nuclear, mitochondrial and chloroplast rRNA genes, group II in mitochondrial and chloroplast mRNA genes. Many of the group I and group II introns are self-splicing in that no additional protein factors are necessary for the intron to be efficiently and accurately excised and the strands reattached.
Group I introns require an external guanosine nucleotide as a cofactor. The 3'-OH of the guanosine nucleotide acts as a nucleophile to attack the 5'-phosphate of the intron's 5' nucleotide. The 3' end of the 5' exon is termed the splice donor site. The 3'-OH at the 3' splice donor end of the 5' exon next attacks the splice acceptor site at the 5' nucleotide of the 3' exon, releasing the intron and covalently attaching the two exons together.
Pre-mRNA processing takes place in the nucleus of eukaryotes, whereas lack of a nuclear membrane in prokaryotes permits initiation of translation while transcription is not yet complete.
Pre-mRNA processing events include capping of the 5’ end on the pre-mRNA, pre-mRNA splicing to remove intronic sequences, and polyadenylation of the 3’ end of the pre-mRNA. animation of RNA splicing requires Flash Player plugin - Download plugin: clickable slide show - spliceosome intron removal :
More in NCBI Molecular Cell Biology on-line text.
proteome
The term proteome refers to an organism’s collection of proteins. A cellular proteome is the collection of proteins found in the particular cell type under a particular set of environmental conditions. The complete proteome for an organism is the potential complete set of proteins, and equates to the sum of cellular proteomes under all possible conditions. The term, introduced in 1995, is analogous to the term genome for nucleic acids, though the proteins themselves are coded for by a portion of the genome.
regulatory proteins
Regulatory proteins bind to segments of DNA and bring about gene gene regulation.
replication
Helicases are a critical part of the DNA replication process because they unwind double-stranded DNA to create single strands suitable for copying by the replication machinery. This and other helicase activity in the cell depends on the ability of the helicase's protein “engine” to crawl along the DNA strand. This locomotion is powered by ATP, the cell's ubiquitous energy source.
Helicase ProteinA helicase protein moving rapidly on a highly flexible single-stranded DNA track. Repetitive movement on the DNA may keep it clear of potentially toxic proteins. Watch Animation 8KB Flash Animation(requires Flash Player)
Helicase ProteinA helicase protein moving rapidly on a highly flexible single-stranded DNA track. Repetitive movement on the DNA may keep it clear of potentially toxic proteins. Watch Animation 8KB Flash Animation(requires Flash Player)
ribosomal structure
MOLECULAR BIOLOGY: ON RNA STRUCTURE
Elucidation of the crystal structures of the ribosome and its subunits have greatly increased the store of information about RNA structure. This information is moving toward an understanding of the principles of RNA folding and of the molecular interactions generating functional capabilities of the ribosome and other RNA systems. Almost all of the possible types of RNA tertiary interactions have been found in ribosomal RNA. One of these interactions is an abundant tertiary structural motif called the A-minor interaction. It has been observed that this participates in both aminoacyl-transfer RNA selection and in peptidyl transferase. It may also play an important role in the structural dynamics of the ribosome.
Elucidation of the crystal structures of the ribosome and its subunits have greatly increased the store of information about RNA structure. This information is moving toward an understanding of the principles of RNA folding and of the molecular interactions generating functional capabilities of the ribosome and other RNA systems. Almost all of the possible types of RNA tertiary interactions have been found in ribosomal RNA. One of these interactions is an abundant tertiary structural motif called the A-minor interaction. It has been observed that this participates in both aminoacyl-transfer RNA selection and in peptidyl transferase. It may also play an important role in the structural dynamics of the ribosome.
RNA polymerase
In prokaryotic cells, all RNA classes are synthesized by a single polymerase. In eukaryotic cells there are 3 distinct classes of RNA polymerase, RNA polymerase (pol) I, II and III. Each polymerase is responsible for the synthesis of a different class of RNA.
RNA polymerase I (Pol I) is dedicated to synthesis of pre-rRNA. RNA polymerase II (Pol II) initiates transcription at DNA sequences corresponding to the 5 Cap of mRNAs and transcribes pre-mRNA. RNA polymerase III (Pol III) transcribes tRNA genes, 5S-rRNA genes, and genes encoding several other small RNAs.
In order to begin transcription, RNA polymerase requires a number of general transcription factors (called TFIIA, TFIIB, and so on). The general transcription factors have been highly conserved in evolution.
All RNA polymerases are dependent upon a DNA template from which to synthesize RNA by addition of complementary bases to an elongating backbone. In RNA, U is substituted for T. The transcribed RNA is complementary to the template strand of the DNA duplex, and is a replica of base-sequences in the non-template, coding strand. The non-template strand is called the coding strand because its sequences are identical to those of the mRNA (with U substituted for T).
p53 - A 3D animation showing the molecule p53 binds to DNA and initiates the transcription of mRNA : diagram - pre-mRNA processing : diagram - intron excision in mRNA precursors : life cycle of an mRNA ~ click on Quicktime Q : alternative splicing - click on fig 1 for animation :
NCBI Molecular Cell Biology - Transcription Initiation Complex : Bacterial Transcription Initiation (NCBI MCB) : SUMMARY transcription initiation (NCBI MCB)
RNA polymerase I (Pol I) is dedicated to synthesis of pre-rRNA. RNA polymerase II (Pol II) initiates transcription at DNA sequences corresponding to the 5 Cap of mRNAs and transcribes pre-mRNA. RNA polymerase III (Pol III) transcribes tRNA genes, 5S-rRNA genes, and genes encoding several other small RNAs.
In order to begin transcription, RNA polymerase requires a number of general transcription factors (called TFIIA, TFIIB, and so on). The general transcription factors have been highly conserved in evolution.
All RNA polymerases are dependent upon a DNA template from which to synthesize RNA by addition of complementary bases to an elongating backbone. In RNA, U is substituted for T. The transcribed RNA is complementary to the template strand of the DNA duplex, and is a replica of base-sequences in the non-template, coding strand. The non-template strand is called the coding strand because its sequences are identical to those of the mRNA (with U substituted for T).
p53 - A 3D animation showing the molecule p53 binds to DNA and initiates the transcription of mRNA : diagram - pre-mRNA processing : diagram - intron excision in mRNA precursors : life cycle of an mRNA ~ click on Quicktime Q : alternative splicing - click on fig 1 for animation :
NCBI Molecular Cell Biology - Transcription Initiation Complex : Bacterial Transcription Initiation (NCBI MCB) : SUMMARY transcription initiation (NCBI MCB)
RNA
In general, a cell has more RNA than DNA, mostly incorporated in ribosomes with a relatively low rate of turnover. Only a small portion of human DNA has coding potential. Almost all RNA sequences in the cytoplasm, being derived from DNA coding sequences, have functional significance, whether transcribed in coding for proteins (mRNA), in performing translation (ribosomal RNA as ribozymes, and tRNA), or in epigenetic mechanisms. Substantial damage to RNA induces apoptosis (cell death), so RNA repair mechanisms are vital to cells.
RNA differs from DNA in its general composition -- ribose is the sugar moiety in the nucleotide backbone, and uracil serves as a base.
RNA differs from DNA in its general composition -- ribose is the sugar moiety in the nucleotide backbone, and uracil serves as a base.
miRNA
microRNAs are a large class of 21 to 24 nucleotide non-coding RNAs that have probable regulatory roles in animals and plants.
mRNA
Messenger RNA, abbreviated mRNA, results from transcription of the base sequence of a segment of single strand archival DNA in the cell's nucleus.
In eukaryotic cells, mRNAs are transcribed as pre-mRNA and processed into mature mRNA in the nucleus, then transported through nuclear pore complexes to the cytoplasm. In some cases, mRNAs are transported to specific areas of the cytoplasm before being translated by ribosomes.
Complex macromolecular machines, such as spliceosomes and the cleavage/ polyadenylation apparatus, control each of these fundamental processes. These macromolecular assemblages comprise scores of proteins, and in many cases RNAs. The complexity of these control-assemblages ensures accuracy in locating promoters and splice sites in the long length of DNA and RNA sequences. Further, the macromolecular machines provide various avenues for regulating synthesis of a polypeptide chain.
Regulation of mRNA stability is central to the post-transcriptional modulation of gene expression. Stability of mRNA varies considerably between mRNAs and can be modulated by extracellular stimuli. Tight control of mRNA stability permits rapid changes mRNA levels, providing a mechanism for prompt termination of protein production. The rate of mRNA decay is determined by cis-acting sequences within the mRNA, which are recognized by trans-acting factors. The best-characterized cis-acting sequences responsible for mRNA decay in mammalian cells are the AU-rich elements, AREs present within the 3'-UTRs of short-lived mRNAs. These AREs are involved in deadenylation and subsequent degradation of mRNAs, and they have also been observed to stimulate 5'-decapping.
Dysregulation of mRNA stability has been associated with different chronic inflammatory diseases, -thalassemia, cancer, and Alzheimer's disease.
A number of ARE binding proteins (ARE-bps) have been identified, which interact with AU- and U-rich regions. These include the ELAV protein family members (most important HuR), the ARE/poly-(U)-binding/degradation factor 1 (AUF-1, also named hnRNP D), the heteronuclear ribonucleoprotein A1 (hnRNP A1), the KH-type splicing regulatory protein (KSRP), tristetraprolin (TTP), the T cell-restricted intracellular antigen (TIA)-1 and the TIA-related protein (TIAR). The KH-type splicing regulatory protein, KSRP has been described as being essential for exosome-mediated degradation of ARE-containing mRNAs.
Full Text Research Article on KSRP-mediated reduction of iNOS expression.
animation - life cycle of an mRNA : animation ~ alternative splicing : NCBI Molecular Cell Biology - Contents : Google mRNA : Classes of RNA Polymerases : Mechanism of RNA Polymerases : Processes of Transcription : Post-transcriptional Processing of RNAs : RNA Splicing : Clinical Significance of Alternative and Aberrant Splicing
In eukaryotic cells, mRNAs are transcribed as pre-mRNA and processed into mature mRNA in the nucleus, then transported through nuclear pore complexes to the cytoplasm. In some cases, mRNAs are transported to specific areas of the cytoplasm before being translated by ribosomes.
Complex macromolecular machines, such as spliceosomes and the cleavage/ polyadenylation apparatus, control each of these fundamental processes. These macromolecular assemblages comprise scores of proteins, and in many cases RNAs. The complexity of these control-assemblages ensures accuracy in locating promoters and splice sites in the long length of DNA and RNA sequences. Further, the macromolecular machines provide various avenues for regulating synthesis of a polypeptide chain.
Regulation of mRNA stability is central to the post-transcriptional modulation of gene expression. Stability of mRNA varies considerably between mRNAs and can be modulated by extracellular stimuli. Tight control of mRNA stability permits rapid changes mRNA levels, providing a mechanism for prompt termination of protein production. The rate of mRNA decay is determined by cis-acting sequences within the mRNA, which are recognized by trans-acting factors. The best-characterized cis-acting sequences responsible for mRNA decay in mammalian cells are the AU-rich elements, AREs present within the 3'-UTRs of short-lived mRNAs. These AREs are involved in deadenylation and subsequent degradation of mRNAs, and they have also been observed to stimulate 5'-decapping.
Dysregulation of mRNA stability has been associated with different chronic inflammatory diseases, -thalassemia, cancer, and Alzheimer's disease.
A number of ARE binding proteins (ARE-bps) have been identified, which interact with AU- and U-rich regions. These include the ELAV protein family members (most important HuR), the ARE/poly-(U)-binding/degradation factor 1 (AUF-1, also named hnRNP D), the heteronuclear ribonucleoprotein A1 (hnRNP A1), the KH-type splicing regulatory protein (KSRP), tristetraprolin (TTP), the T cell-restricted intracellular antigen (TIA)-1 and the TIA-related protein (TIAR). The KH-type splicing regulatory protein, KSRP has been described as being essential for exosome-mediated degradation of ARE-containing mRNAs.
Full Text Research Article on KSRP-mediated reduction of iNOS expression.
animation - life cycle of an mRNA : animation ~ alternative splicing : NCBI Molecular Cell Biology - Contents : Google mRNA : Classes of RNA Polymerases : Mechanism of RNA Polymerases : Processes of Transcription : Post-transcriptional Processing of RNAs : RNA Splicing : Clinical Significance of Alternative and Aberrant Splicing
tRNA
Largest Computational Biology Simulation Mimics Life's Most Essential Nanomachine: "The simulations also reveal that the essential translating molecule, transfer RNA, must be flexible in two places for decoding to occur, furthering the growing belief that transfer RNA is a major player in the machine-like movement of the ribosome. The simulation also sets the stage for future biochemical research into decoding by identifying 20 universally conserved ribosomal bases important for accommodation, as well as a new structural gate, which may act as a control mechanism during transfer RNA selection." The original news release can be found here. : Voxel simulation image : Image 2 : Image 3 : Image 4 : Image 5 : High res movie : Low res movie : See an image of the Q machine at http://www.lanl.gov/asci/.
ribozymes in repair of RNA and DNA
Ribozyme-mediated revision of RNA and DNA -- Long et al. 112 (3): 312 -- Journal of Clinical Investigation: "Several classes of catalytic RNAs, or ribozymes, have the capacity to revise genetic information. Group I and Group II ribozymes are derived from naturally occurring Group I and Group II introns, respectively. These introns are found in the genes of a variety of lower eukaryotes and prokaryotes. They differ fundamentally from spliceosomal introns since Group I and Group II introns self-splice from the precursor RNA independent of the spliceosome. The intron adopts the catalytic structure that is capable of cleaving RNA splice sites and ligating the flanking exons together. In addition to self-splicing from RNA precursors, some Group II introns are able to reverse-splice into DNA. The splicing activity of these introns can be harnessed as a molecular tool that may potentially revise any gene of interest."
BIO.COM: Biotechnology & Pharmaceutical News, Jobs, Software, Reports, Books, Events: "Hairpin ribozyme . . . minimal RNA enzyme was discovered by botanists in plants infected by tobacco ring spot virus and a number of other viruses. It is what is known as plant satellite RNA.Satellite RNAs are parasitic pieces of RNA that are not exactly viruses because they don't encode for proteins. Instead, they catalyze simple cut-and-paste reactions in order to replicate themselves, exacerbating or ameliorating diseases caused by plant viruses. A plant that has tobacco ring spot virus, for instance, will be more diseased if it also has this satellite RNA. "
BIO.COM: Biotechnology & Pharmaceutical News, Jobs, Software, Reports, Books, Events: "Hairpin ribozyme . . . minimal RNA enzyme was discovered by botanists in plants infected by tobacco ring spot virus and a number of other viruses. It is what is known as plant satellite RNA.Satellite RNAs are parasitic pieces of RNA that are not exactly viruses because they don't encode for proteins. Instead, they catalyze simple cut-and-paste reactions in order to replicate themselves, exacerbating or ameliorating diseases caused by plant viruses. A plant that has tobacco ring spot virus, for instance, will be more diseased if it also has this satellite RNA. "
silencer
Silencers are control sections of DNA that, like enhancers, may be located thousands of base pairs away from the gene they regulate. However, when transcription factors bind to them, expression of the gene they regulate is repressed. This is the opposite effect to that of enhancers, which increase rate of transcription.
"One basic premise of chromatin regulation is that genes are silenced through compaction of chromatin, which reduces the accessibility of DNA. In contrast, gene expression may require the "opening up" of chromatin. The Polycomb group (PcG) of gene repressors and the trithorax group (trxG) of gene activators are two antagonistic classes of proteins that may act through modulation of chromatin structure. Together, these factors maintain the gene expression patterns of key developmental regulators and hence are crucial players in cellular differentiation, stem cell renewal, and cancer." in:
MOLECULAR BIOLOGY: CHROMATIN DNA PACKAGING AND GENE SILENCING
"One basic premise of chromatin regulation is that genes are silenced through compaction of chromatin, which reduces the accessibility of DNA. In contrast, gene expression may require the "opening up" of chromatin. The Polycomb group (PcG) of gene repressors and the trithorax group (trxG) of gene activators are two antagonistic classes of proteins that may act through modulation of chromatin structure. Together, these factors maintain the gene expression patterns of key developmental regulators and hence are crucial players in cellular differentiation, stem cell renewal, and cancer." in:
MOLECULAR BIOLOGY: CHROMATIN DNA PACKAGING AND GENE SILENCING
spliceosome
Almost all eukaryotic protein-coding genomes contain non-coding intervening sequences called introns. Spliceosomes are complex ribonuclear machines in eukaryotes that remove the non-coding introns from primary transcript, precursor mRNA (pre-mRNA or hnRNA). Alternatively, the term spliceome can be used to describe the complete set of all possible alternative splices in an organism, analogous to the genome or proteome.
Spliceosomes are variably composed of as many as 300 distinct proteins and five RNAs, making them among the most complex macromolecular machines known. review article abstract. Essential components of the spliceosome are the small RNA-protein complexes called small nuclear ribonucleoproteins (snRNPs, pronounced 'snurps'). These are U1, U2, U4, U5, and U6, so named because they are rich in uridine nucleotides. In addition to snRNPs, splicing requires many non-snRNP protein factors. The snRNPs participate in several RNA-RNA and RNA-protein interactions.
The spliceosome recognizes specific 5' and 3' sites on the pre-mRNA. The intronal area between these locations is excised, and the two exons are spliced (ligated). diagram - formation of a spliceosome : diagram - intron excision in mRNA precursors : diagram - pre-mRNA processing : animation ~ alternative splicing : animation of RNA splicing requires Flash Player plugin - Download plugin: clickable slide show - spliceosome intron removal : alternative splicing - click on fig 1 for animation : life cycle of an mRNA ~ click on Quicktime Q : clickable slide show - spliceosome intron removal :
Aberrant splicing creates mutant proteins, while alternative splicing generates diversity. Splice variants and epigenetic mechanisms account for the ability of about 25,000 human genes to code for about 100,000 human proteins. In addition to this variation, and that provided by recombination, each human gene possesses at least two isoforms – one from each parent. X Inactivation
Spliceosomes are variably composed of as many as 300 distinct proteins and five RNAs, making them among the most complex macromolecular machines known. review article abstract. Essential components of the spliceosome are the small RNA-protein complexes called small nuclear ribonucleoproteins (snRNPs, pronounced 'snurps'). These are U1, U2, U4, U5, and U6, so named because they are rich in uridine nucleotides. In addition to snRNPs, splicing requires many non-snRNP protein factors. The snRNPs participate in several RNA-RNA and RNA-protein interactions.
The spliceosome recognizes specific 5' and 3' sites on the pre-mRNA. The intronal area between these locations is excised, and the two exons are spliced (ligated). diagram - formation of a spliceosome : diagram - intron excision in mRNA precursors : diagram - pre-mRNA processing : animation ~ alternative splicing : animation of RNA splicing requires Flash Player plugin - Download plugin: clickable slide show - spliceosome intron removal : alternative splicing - click on fig 1 for animation : life cycle of an mRNA ~ click on Quicktime Q : clickable slide show - spliceosome intron removal :
Aberrant splicing creates mutant proteins, while alternative splicing generates diversity. Splice variants and epigenetic mechanisms account for the ability of about 25,000 human genes to code for about 100,000 human proteins. In addition to this variation, and that provided by recombination, each human gene possesses at least two isoforms – one from each parent. X Inactivation
SSOs
Increased efficiency of oligonucleotide-mediated gene repair through slowing replication fork progression -- Wu et al. 102 (7): 2508 -- Proceedings of the National Academy of Sciences: "In mammalian cells, the precise mechanism of SSO-mediated chromosome alteration remains to be established, and there have been problems in obtaining reproducible targeting efficiencies. It has previously been suggested that the chromatin structure, which changes throughout the cell cycle, may be a key factor underlying these variations in efficiency. "
trans-splicing ribozymes and therapeutics
Ribozyme-mediated revision of RNA and DNA -- Long et al. 112 (3): 312 -- Journal of Clinical Investigation: "The therapeutic revision of RNA by ribozymes is mechanistically different from that of DNA. To revise RNA sequences, a Group I intron from Tetrahymena thermophila has been adapted for trans-splicing. Trans-splicing ribozymes splice therapeutic RNA sequences onto a target transcript in a process called ribozyme-mediated RNA repair. Considerable progress has been made in the development of trans-splicing ribozymes for therapeutic applications such as treating sickle cell disease and cancer." This is one of the relatively recent targeted genetic repair techniques.
"The spliceosome has also been shown to trans-splice RNA transcripts. Spliceosomal-mediated RNA trans-splicing is also called called SMaRT."
"The spliceosome has also been shown to trans-splice RNA transcripts. Spliceosomal-mediated RNA trans-splicing is also called called SMaRT."
transcription
In transcription, an RNA polymerase enzyme (RNAp, or pol III in eukaryotes) directs generation of a complementary strand of mRNA from DNA. The mechanism of alternative splicing enables to production of different mature mRNA molecules, depending on what sequences are treated as introns and what remain as exons.
Transcription involves 3 phases: initiation, elongation, and termination.
1. RNA polymerase II initiates transcription at the first nucleotide of the first exon of a gene.
2. The 5 end of the nascent RNA is capped with 7-methylguanylate (capping).
3. Transcription by RNA polymerase II terminates at any one of multiple termination sites downstream from the poly(A) site, which is located at the 3 end of the final exon.
4. After the primary transcript is cleaved at the poly(A) site.
5. A string of adenine (A) residues is added (polyadenylation). The poly(A) tail contains ≈250 A residues in mammals, ≈150 in insects, and ≈100 in yeasts.
In more detail: RNA polymerase binds to the promoter region of one strand of DNA (5’ end), and the DNA double helix is un-zipped into single strands. First, RNA polymerase requires a number of general transcription factors (called TFIIA, TFIIB, etc.). The promoter contains a DNA sequence called the TATA box, which is located 25 nucleotides away from the site of initiation of transcription. The TATA box is recognized and bound by transcription factor TFIID, which then enables the adjacent binding of TFIIB. The rest of the general transcription factors plus the RNA polymerase assemble at the promoter. diagram - initiation of transcription.
The RNAp enzyme moves toward the 3’ end, connecting complementary bases into an elongating chain of RNA nucleotides. At termination, the transcribed mRNA molecule is released from the DNA strand. In prokaryotic cells – without a nuclear membrane – translation may begin prior to termination. In eukaryotic cells – with a nuclear membrane – the processed mRNA moves through the nuclear pores into the cytoplasm, where ribosomes on the rough endoplasmic reticulum translate the mRNA code into a peptide or a protein. Epigenetic, alternative splicing mechanisms can edit the mRNA prior to its translation into protein.
Capping of the 5’ end on the pre-mRNA with 7-methylguanylate occurs soon after initiation of transcription, and the 5 cap is retained in mature mRNAs.
Cleavage, pre-mRNA splicing, and polyadenylation usually follow termination of transcription of short primary transcripts with few introns. However, introns often are spliced out of the nascent RNA before transcription of the gene is complete for large genes with multiple introns.
It was believed that most genes in higher eukaryotes are regulated by controlling their transcription. However, it is increasingly recognized that epigenetic mechanisms (such as alternative splicing) are important in generating many proteins from a single gene, accounting for the Human Genome Project’s discovery that a mere 30,000 genes code for about 100,000 proteins.
animation - start of transcription : animation - life cycle of an mRNA : animation ~ alternative splicing
More at NCBI Molecular Cell Biology - Transcription Initiation Complex : SUMMARY transcription initiation (NCBI MCB) : Processes of Transcription : Wikipedia : Central Dogma :
Transcription involves 3 phases: initiation, elongation, and termination.
1. RNA polymerase II initiates transcription at the first nucleotide of the first exon of a gene.
2. The 5 end of the nascent RNA is capped with 7-methylguanylate (capping).
3. Transcription by RNA polymerase II terminates at any one of multiple termination sites downstream from the poly(A) site, which is located at the 3 end of the final exon.
4. After the primary transcript is cleaved at the poly(A) site.
5. A string of adenine (A) residues is added (polyadenylation). The poly(A) tail contains ≈250 A residues in mammals, ≈150 in insects, and ≈100 in yeasts.
In more detail: RNA polymerase binds to the promoter region of one strand of DNA (5’ end), and the DNA double helix is un-zipped into single strands. First, RNA polymerase requires a number of general transcription factors (called TFIIA, TFIIB, etc.). The promoter contains a DNA sequence called the TATA box, which is located 25 nucleotides away from the site of initiation of transcription. The TATA box is recognized and bound by transcription factor TFIID, which then enables the adjacent binding of TFIIB. The rest of the general transcription factors plus the RNA polymerase assemble at the promoter. diagram - initiation of transcription.
The RNAp enzyme moves toward the 3’ end, connecting complementary bases into an elongating chain of RNA nucleotides. At termination, the transcribed mRNA molecule is released from the DNA strand. In prokaryotic cells – without a nuclear membrane – translation may begin prior to termination. In eukaryotic cells – with a nuclear membrane – the processed mRNA moves through the nuclear pores into the cytoplasm, where ribosomes on the rough endoplasmic reticulum translate the mRNA code into a peptide or a protein. Epigenetic, alternative splicing mechanisms can edit the mRNA prior to its translation into protein.
Capping of the 5’ end on the pre-mRNA with 7-methylguanylate occurs soon after initiation of transcription, and the 5 cap is retained in mature mRNAs.
Cleavage, pre-mRNA splicing, and polyadenylation usually follow termination of transcription of short primary transcripts with few introns. However, introns often are spliced out of the nascent RNA before transcription of the gene is complete for large genes with multiple introns.
It was believed that most genes in higher eukaryotes are regulated by controlling their transcription. However, it is increasingly recognized that epigenetic mechanisms (such as alternative splicing) are important in generating many proteins from a single gene, accounting for the Human Genome Project’s discovery that a mere 30,000 genes code for about 100,000 proteins.
animation - start of transcription : animation - life cycle of an mRNA : animation ~ alternative splicing
More at NCBI Molecular Cell Biology - Transcription Initiation Complex : SUMMARY transcription initiation (NCBI MCB) : Processes of Transcription : Wikipedia : Central Dogma :
termination of transcription
MOLECULAR BIOLOGY: ON TRANSCRIPTION TERMINATION: "Transcription termination is important, not least because stopping too late will disrupt the regulation of other genes on the same chromosome; stopping prematurely, meanwhile, would produce truncated and therefore defective mRNAs. . . termination on most genes has been found to occur at various positions rather than at a single site. Moreover, a discrete signal in the DNA that might define the termination site could not be identified. . . Recent work(1-3) address this problem and provides evidence in favor of a striking alternative mechanism: that the transcribing RNA polymerase is "torpedoed". The mRNA is cut while it is still being synthesized, with the liberated region forming the mRNA. The remaining RNA trails out of the transcribing RNA polymerase, and is attacked by an exonuclease -- an enzyme that can degrade the RNA from its free end. This enzyme chases after the polymerase, chewing up the RNA strand as it goes. When it catches up with the polymerase, transcription is terminated."